Research
Research Mission
I co-lead the KERMIT research unit together with Bernard De Baets, where I am responsible for the computational modelling research line. KERMIT’s mission is to harness mathematics and computation to unravel life’s complexities, optimize biological functions, and drive innovation in biodesign and decision-making under uncertainty.
In our subgroup, we focus on non-conventional computing for understanding biological systems, including stochastic and differential programming, evolutionary computing, and other related approaches. Though we have an interest in various application fields within the applied biological sciences, we have plant growth and synthetic biology as our key focuses.
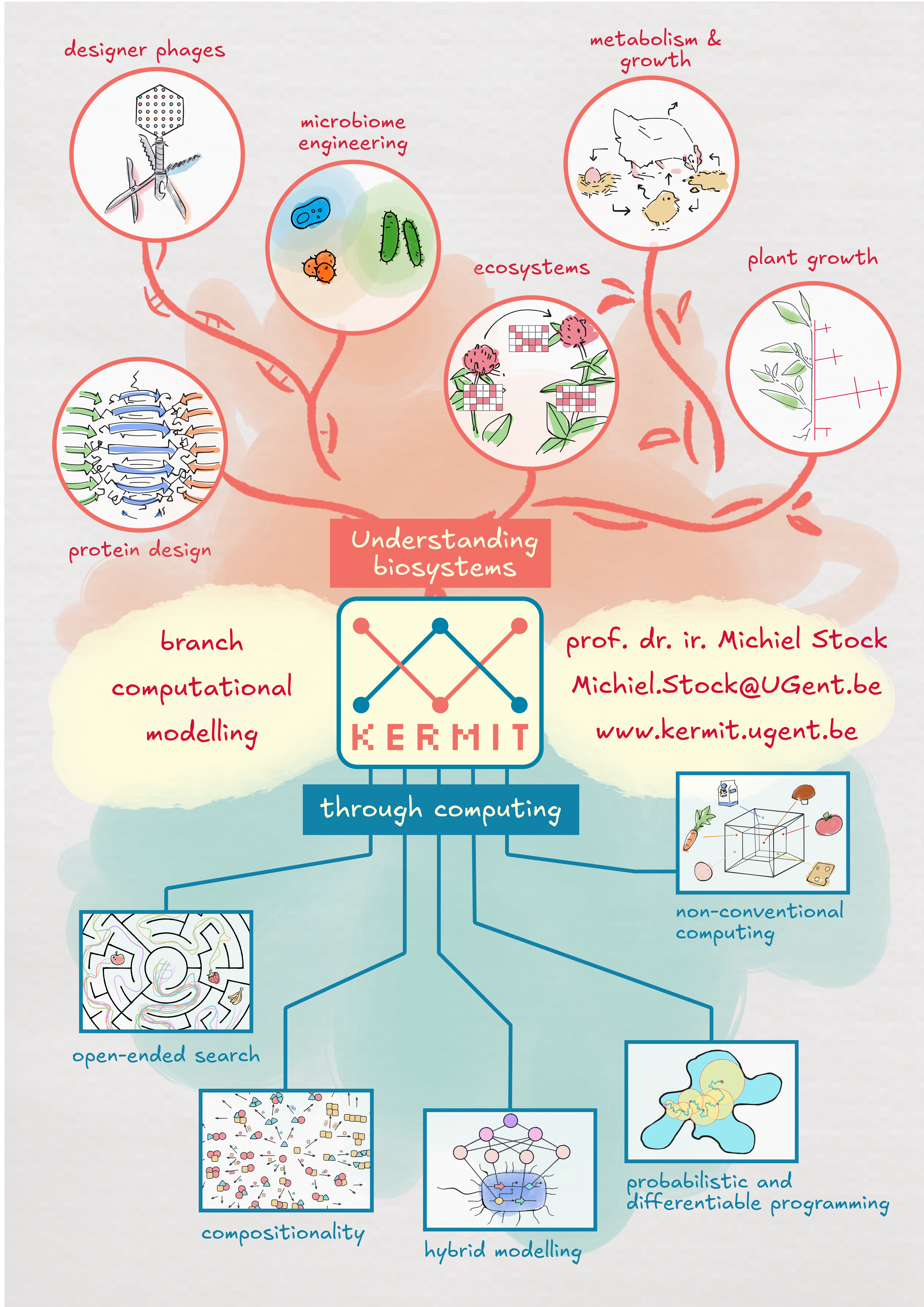
Research Themes
Computational Modelling for Synthetic Biology
We develop AI-driven methods to design and optimize biological parts and systems. This includes predicting promoter strength, designing proteins, and exploring the open-ended nature of biological innovation.
Machine Learning for Ecological Networks
Using optimal transportation theory and maximum entropy principles, we model how species interact in ecological networks — predicting pollination, host-parasite, and food web patterns.
Hyperdimensional Computing for Biological Data
We pioneer the use of hyperdimensional computing as a fast, robust, and interpretable paradigm for analyzing biological sequences, molecular data, and ecological patterns.
Plant Growth Modelling
Combining differential equations, sensor data, and machine learning to model and predict plant growth dynamics under varying environmental conditions.
Quality-Diversity & Evolutionary Algorithms
We adapt quality-diversity algorithms from evolutionary computation to discover diverse, high-performing solutions in biological design and optimization problems.
Key Publications
Open-endedness in synthetic biology: A route to continual innovation for biological design
Science Advances, 10, eadi3621 (2024)
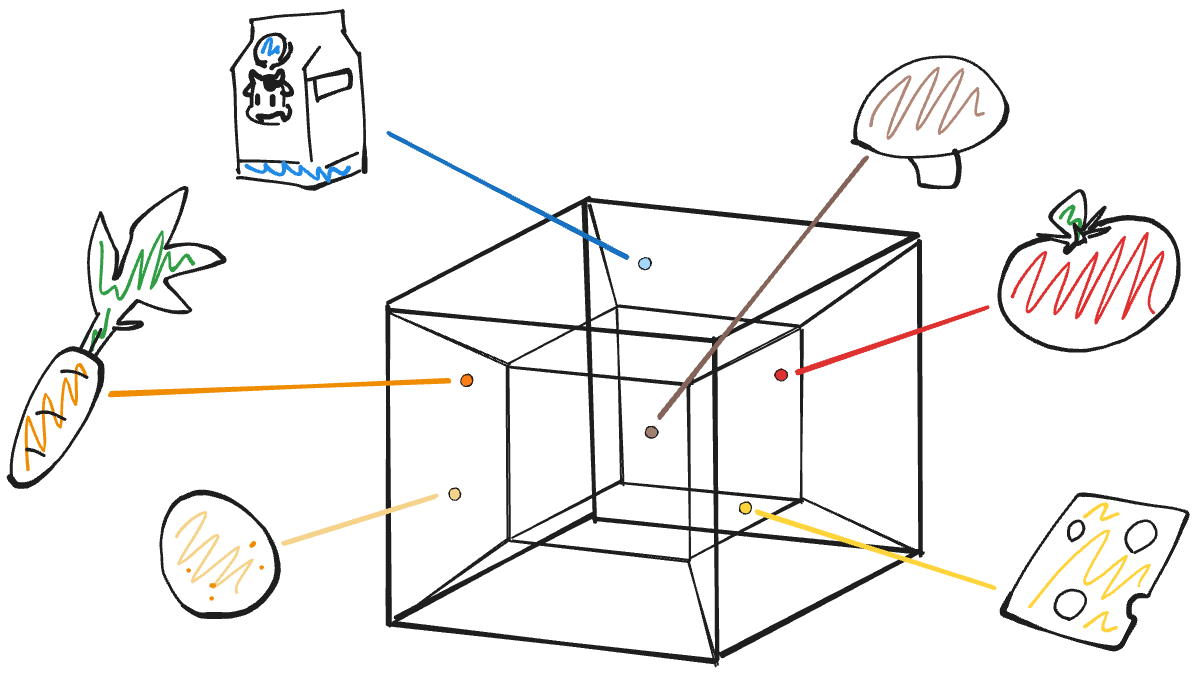
Hyperdimensional computing: a fast, robust and interpretable paradigm for biological data
PLOS Computational Biology, 20, e1012426 (2024)
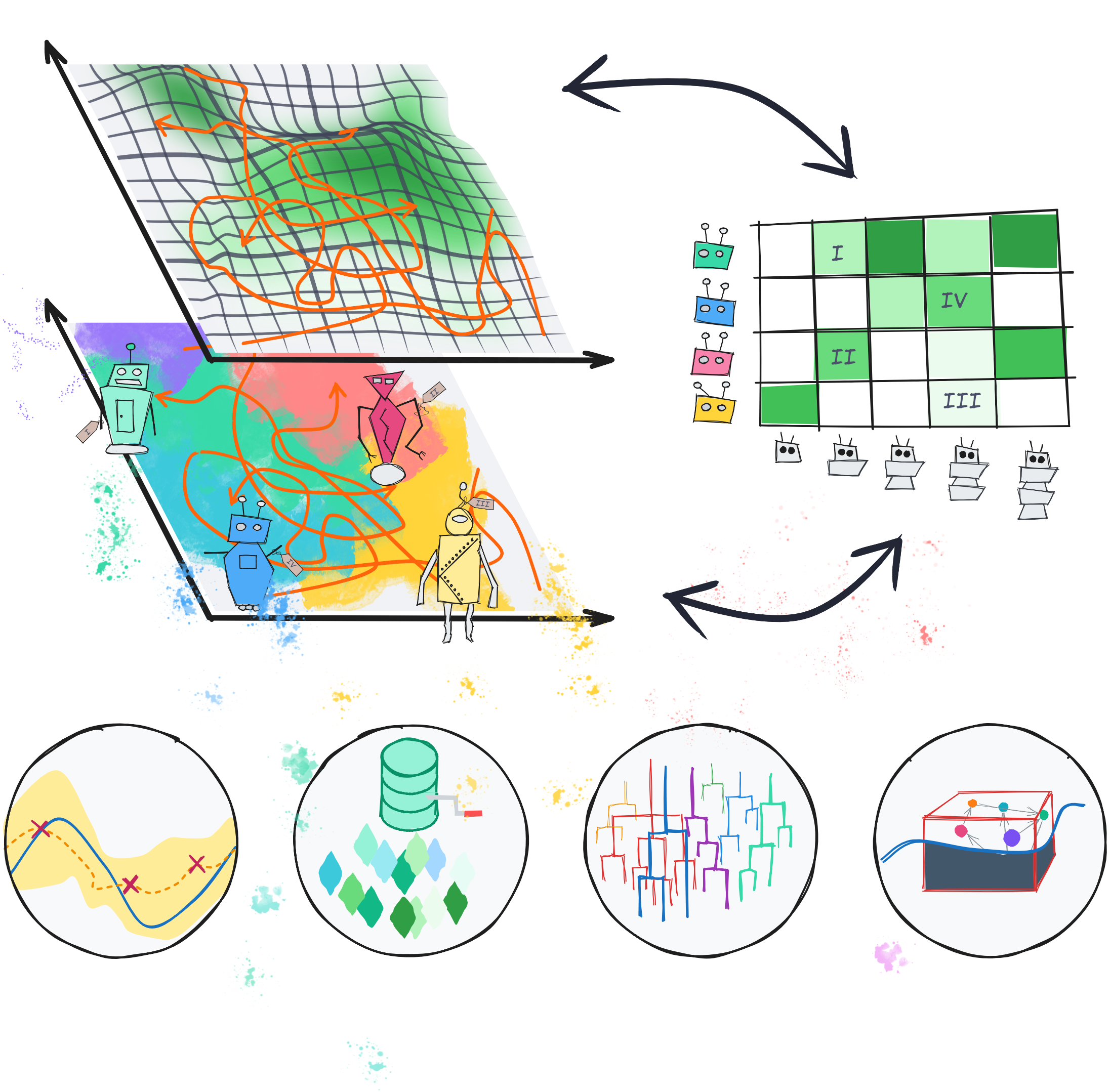
Quality-diversity methods for the modern data scientist
WIREs Computational Statistics, 17, e70047 (2025)
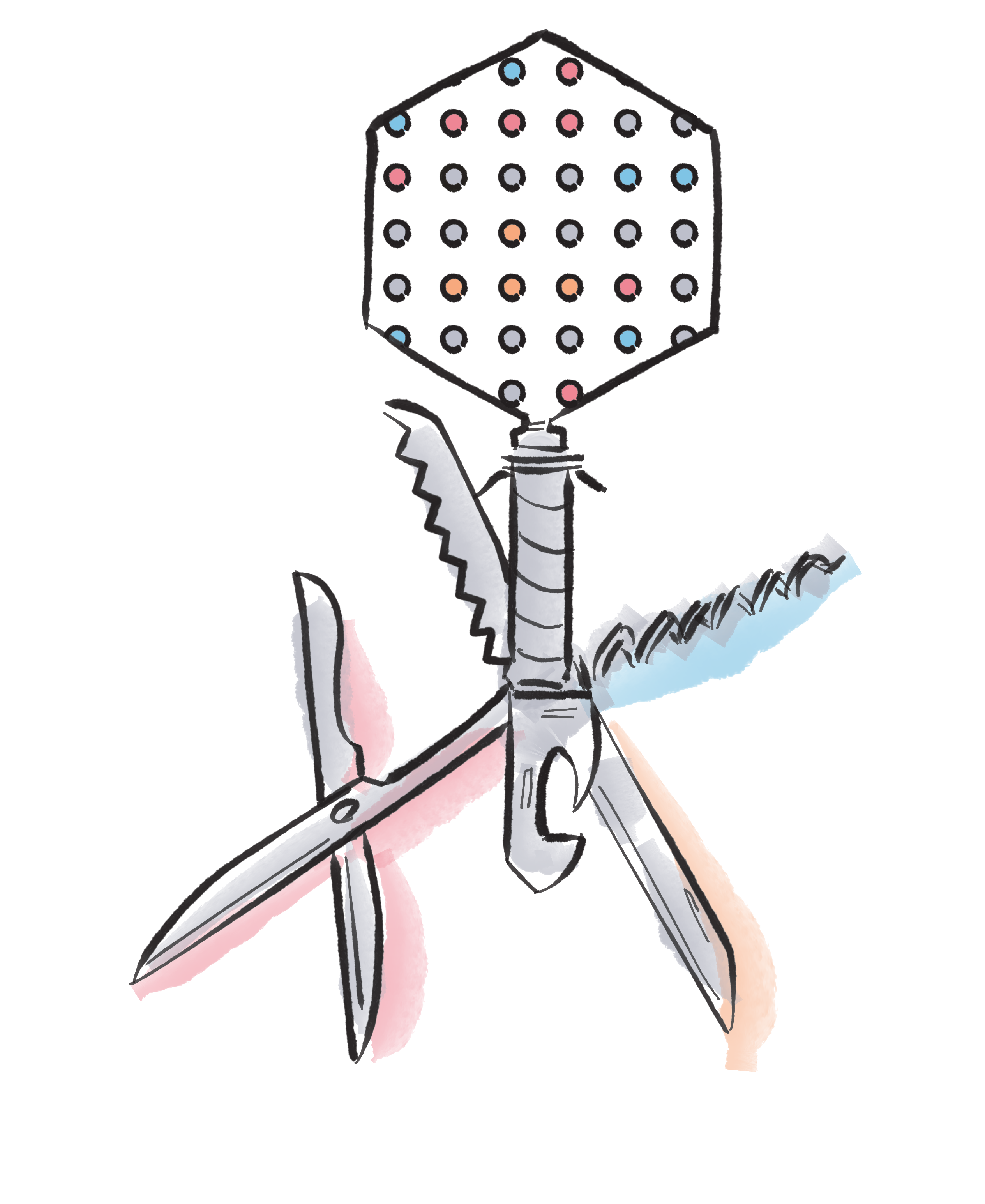

Prediction of Klebsiella phage-host specificity at the strain level
Nature Communications, 15, 4355 (2024)
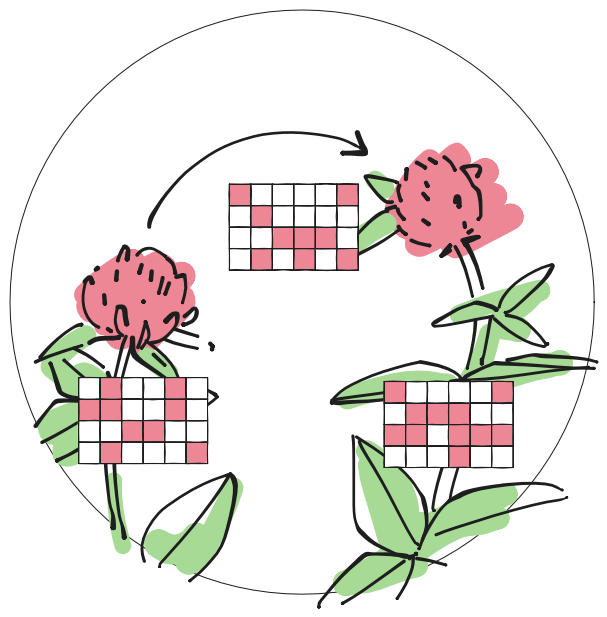

Optimal transportation theory for species interaction networks
Ecology and Evolution, 11, 3841–3855 (2021)

Plant science in the age of simulation intelligence
Frontiers in Plant Science, 14, 1299208 (2024)
For a complete list, see Google Scholar or the UGent Academic Bibliography.
Open Positions
If you are interested in a PhD or postdoc position in computational modelling for biological systems, please get in touch.
I also supervise MSc thesis projects — see the Teaching page.